GENOMIC
Mapping
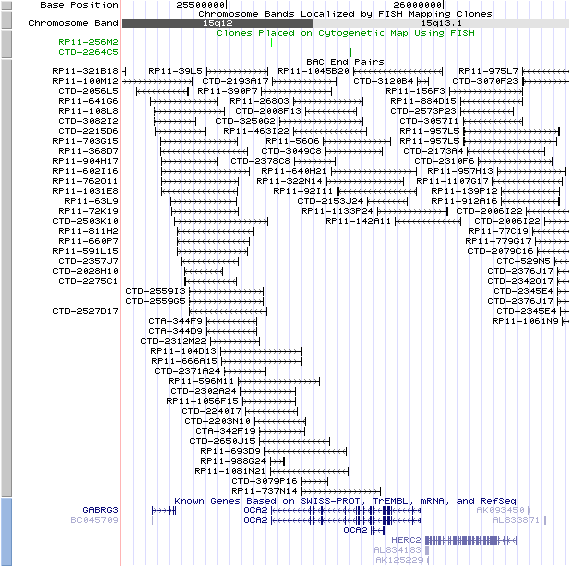
15q13.1 View the map and BAC clones (data from UCSC genome browser).
Note that the human P gene corresponds to the region within the chromosome segment 15q11-q13, which is typically deleted in patients with Prader-Willi and Angelman syndrome.
Structure
(assembly 07/03)
OCA2/NM_000275: 24 exons, 344,434bp, Chr15: 25,602,392 - 25,946,825.
The figure below shows the structure of the OCA2 gene (data from UCSC genome browser).
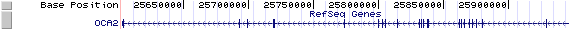
Regulatory Element
Search the 5'UTR and 1kb upstream regions (human and mouse) by CONREAL with 80% Position Weight Matrices (PWMs) threshold (view results here).
TRANSCRIPT
RefSeq/ORF
OCA2(NM_000275), 3,136bp, view ORF and the alignment to genomic.
Expression Pattern
Tissue specificity: Varied expression, highest in brain.
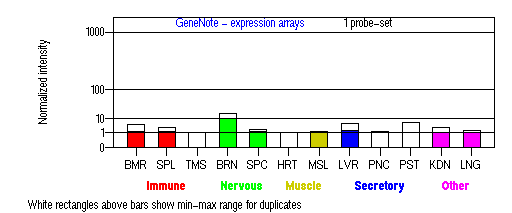
BMR: Bone marrow; SPL: Spleen; TMS: Thymus; BRN: Brain; SPC: Spinal cord; HRT: Heart; MSL: Skeletal muscle;
LVR; Liver; PNC: Pancreas; PST: Prostate; KDN: Kidney; LNG: Lung. (data from GeneCards )
PROTEIN
Sequence
P protein (NP_000266): 838aa, ExPaSy NiceProt view of Swiss-Prot:Q04671.
Synonyms: pink-eye dilution (murine) homolog; Melanocyte-specific transporter protein.
Ortholog
| Species | Mouse | Zebrafish | Fruitfly | Mosquito |
| GeneView | p | 07116 | CG15624/hoe2 | 1280250 |
| Protein | NP_068679 (833aa) | 13636 (128aa) | NP_608878 (846aa) | XP_320080 (580aa) |
| Identities | 78% /659aa | 59% /76aa | 40% /232aa | 51% /266aa |
View multiple sequence alignment (PDF file) by ClustalW and GeneDoc.
Domain
(1) Domains predicted by SMART:
a) low complexity 123 - 135
b) transmembrane 175 - 197
c) transmembrane 327 - 346
d) transmembrane 353 - 370
e) transmembrane 385 - 407
f) transmembrane 420 - 439
g) transmembrane 508 - 530
h) low complexity 580 - 591
i) transmembrane 621 - 643
j) transmembrane 648 - 667
k) transmembrane 679 - 701
l) transmembrane 721 - 739
m) transmembrane 777 - 799
n) transmembrane 814 - 836
(2) Transmembrane domains predicted by SOSUI: 12 transmembrane helices detected.
| No. | N terminal | transmembrane region | C terminal | type | length |
| 1 | 176 | QWLKVMGLFAFVVLCSILFSLYP | 198 | PRIMARY | 23 |
| 2 | 326 | SVETQVTIATAILAGVYALIIFE | 348 | PRIMARY | 23 |
| 3 | 352 | RTLAAMLGSLAALAALAVIGDRP | 374 | PRIMARY | 23 |
| 4 | 384 | DFETLALLFGMMILVAIFSETGF | 406 | PRIMARY | 23 |
| 5 | 423 | WAMIIMLCLIAAVLSAFLDNVTT | 445 | PRIMARY | 23 |
| 6 | 501 | MGLDFAGFTAHMFIGICLVLLVC | 523 | SECONDARY | 23 |
| 7 | 619 | DGILLAKCLTVLGFVIFMFFLNS | 641 | PRIMARY | 23 |
| 8 | 648 | LDLGWIAILGAIWLLILADIHDF | 670 | PRIMARY | 23 |
| 9 | 679 | WATLLFFAALFVLMEALAHLHLI | 701 | PRIMARY | 23 |
| 10 | 708 | TALLIKMVPEEQRLIAAIVLVVW | 730 | SECONDARY | 23 |
| 11 | 769 | MYALAFGACLGGNGTLIGASANV | 791 | SECONDARY | 23 |
| 12 | 811 | RLGFPMMVVSCTVGMCYLLVAHV | 833 | PRIMARY | 23 |
(3) Graphical view of InterPro domain structure.
Motif/Site
(1) Predicted results by ScanProsite:
a) Amidation site : [occurs frequently]
7 - 10: dGRR,
33 - 36: aGKR.
b) Protein kinase C phosphorylation site : [occurs frequently]
86 - 88: SsR,
153 - 155: SeK,
303 - 305: SiR,
453 - 455: TiR.
c) N-glycosylation site : [occurs frequently]
214 - 217: NYSV,
218 - 221: NLSS,
273 - 276: NWTV,
442 - 445: NVTT,
754 - 757: NLSH,
781 - 784: NGTL.
(2) Predicted results of subprograms by PSORT II:
a) Seems to have no N-terminal signal peptide
b) KDEL ER retention motif in the C-terminus: none
c) ER Membrane Retention Signals: none
d) VAC possible vacuolar targeting motif: none
e) Actinin-type actin-binding motif: type 1: none; type 2: none
f) Prenylation motif: none
g) memYQRL transport motif from cell surface to Golgi: none
h) Tyrosines in the tail: none
i) Dileucine motif in the tail: none
3D Model
(1) ModBase entry found, results here.
(2)ModBase Predicted Comparative 3D Structure of Q04671 from UCSC Genome Sorter.
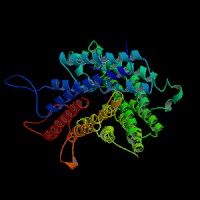
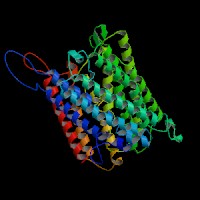
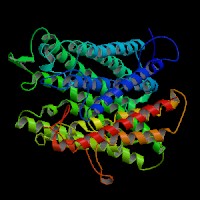
From left to right: Front, Top, and Side views of predicted protein
2D-PAGE
This protein does not exist in the current release of SWISS-2DPAGE.
Computed theoretical MW=92,894Da, pI=6.72 (NP_000266).
FUNCTION
Ontology
a) Biological process: eye pigment biosynthesis
b) L-tyrosine, eye pigment precursor, and arsenite transporter activity
c) Serine-type endopeptidase inhibitor activity
d) Integral to membrane
e) Belongs to the SLC13A family of transporters (P subfamily).
Location
Integral membrane protein. Steady-state melanosomal localization requires a conserved consensus acidic dileucine-based sorting motif within the cytoplasmic N-terminal region of OCA2. A second dileucine signal within this region confers steady-state lysosomal localization in melanocytes (Sitaram, et al).
Interaction
The P protein regulates posttranslational processing and transport of tyrosinase (Chen, et al; Toyofuku, et al).
No entries in the CuraGen Drosophila interaction database for CG15624.
Pathway
The P protein is believed to be an integral membrane protein involved in small molecule transport, specifically tyrosine - a precursor of melanin. P protein plays an important role in the generation or maintenance of melanosomal pH, which is required for the tyrosinase activity and the assembly of the normal melanogenic complex (Puri, et al). P protein increases cellular sensitivity to arsenicals and other metalloids and can modulate intracellular glutathione metabolism (Staleva, et al).
MUTATION
Allele or SNP
72 mutational alleles deposited in HGMD.
SNPs deposited in dbSNP.
Selected allelic examples described in OMIM.
Distribution
The distribution of mutations and polymorphisms is described in detail in Albinism Database. For cross-reference, view the distribution of the same set of mutations and polymorphisms in the Retina International Mutation Database of P-Gene.
Except the mutations deposited in the HGMD database, 44 additional mutations are listed here.
(1) Duan, et al and Li, et al have reported four novel mutations (N476D, G775R, A787T, Y827H) in the P gene in Chinese OCA2 patients.
(2) Three mutations in P (V443I, N476S, C793F) were identified by Preising, et al.
(3) An Thai-Chinese OCA2 patient was compound heterozygous for two novel splice site mutations, a paternally inherited IVS9 + 1delG, and a maternally inherited IVS23 +
1G >A (Wattanasirichaigoon, et al).
(4) Hutton, et al (2008) reported one novel mutation (I634N) in a non-Hispanic Caucasian OCA2 patient.
(5) Dai, et al reported one novel mutations (T450M) in the P gene in Chinese OCA2 patients.
(6) Rooryck, et al (2008) reported 12 novel mutations including six missense (H249D, G485V, A558P, S732L, P743R, R811S), two frameshift (c.898insG, c.2106delinsTTC), one splicing (c. 1116 +6T>C), one nonsense
mutation (W61X), and two deletions spanning exons 3-20.
(7) Gronskov, et al (2009) reported 6 novel mutations (c.157delA, E96A, Y342C, Y415H, C626R, L727P) in Danish OCA or AROA patients.
(8) Wei, et al (2010) reported 14 previously unreported
mutational alleles (c.168delC, c.980insT, c.860_883del24 bp, c.2351_2376del26 bp, p.R136X, p.K155N, p.R243C, p.Q321P, p.C430X, p.T450K, p.R455G, p.R555C, p.A776D, and p.G782R) in Chinese Han OCA patients. The most frequent allele in this screen was p.R455G, which accounts for 15% of the mutational OCA2 alleles in this population.
(9) Johanson, et al (2010) identified one novel mutation (p.G775D) in Polynesian OCA2 patients.
Effect
Most missense mutations of the p gene are located within the loops between the TM domains. Genotype-phenotype relationship is not well defined.
PHENOTYPE
Defects in OCA2 are the cause of oculocutaneous albinism type II (OCA2) (Rinchik, et al) (OMIM 203200). OCA2 is an autosomal recessive form of albinism, a disorder of pigmentation in the skin, hair, and eyes. The phenotype of patients with OCA2 is typically somewhat less severe than in those with tyrosinase-deficient OCA1. There are several forms of OCA2, from typical OCA to relatively mild 'autosomal recessive ocular albinism' (AROA) (OMIM 203310). OCA2 belongs to tyrosinase-positive oculocutaneous albinism. OCA2 patients are usually born with minimal pigmentation and become darker during adulthood, finally showing minimal-to-moderate pigmentation of their skin, hair and eyes.
OCA2 is about twice as common as OCA1 in African and African-American populations. The frequency of OCA2 is greatly increased in patients with Prader-Willi or Angelman syndrome (Lee, et al). Recent studies have shown that OCA2 is not the most frequent OCA subtype in some populations. In Chinese, Japanese and Caucasian patients, the most prevalent form of OCA is OCA1, whereas OCA2 and OCA4 are much less frequent and OCA3 virtually non-existent (Hutton, et al (2008); Suzuki, et al (2008); Rooryck, et al (2008); Gronskov, et al (2009); Wei, et al (2010) ). Genome-wide association studies have shown that OCA2 is associated with the colors of human skin, hair and iris (Han, et al). The non-synonymous polymorphism rs1800414 (His615Arg) located within the OCA2 gene is significantly associated with skin pigmentation in East Asian populations (Edwards, et al).
REFERENCE
- Chen K, Manga P, Orlow SJ. Pink-eyed dilution protein controls the processing of tyrosinase. Mol Biol Cell 2002; 13: 1953-64. PMID: 12058062
- Dai C, Li W, Gao B, Li L, Lu G. Mutation screening of the TYR and P gene in three patients with oculocutaneous albinism. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2008; 25: 373-377. PMID: 18683130
- Duan HL, Li HY, Wu WQ, Zheng H, Chen Z. A novel P gene mutation in a Chinese family with oculocutaneous albinism. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 2006; 23: 614-7. PMID: 17160937
- Edwards M, Bigham A, Tan J, Li S, Gozdzik A, Ross K, Jin L, Parra EJ. Association of the OCA2 Polymorphism His615Arg with Melanin Content in East Asian Populations: Further Evidence of Convergent Evolution of Skin Pigmentation. PLoS Genet 2010; 6: e1000867.PMID: 20221248
- Gronskov K, Ek J, Sand A, Scheller R, Bygum A, Brixen K, Brondum-Nielsen K, Rosenberg T. Birth prevalence and mutation spectrum in Danish patients with autosomal recessive albinism. Invest Ophthalmol Vis Sci 2009; 50: 1058-64. PMID: 19060277
- Han J, Kraft P, Nan H, Guo Q, Chen C, Qureshi A, Hankinson SE, Hu FB, Duffy DL, Zhao ZZ, Martin NG, Montgomery GW, Hayward NK, Thomas G, Hoover RN, Chanock S, Hunter DJ. A genome-wide association study identifies novel alleles associated with hair color and skin pigmentation. PLoS Genet 2008; 4:e1000074. PMID: 18483556
- Hutton SM, Spritz RA. Comprehensive Analysis of Oculocutaneous Albinism among Non-Hispanic Caucasians Shows that OCA1 Is the Most Prevalent OCA Type. J Invest Dermatol 2008; 128: 2442-50. PMID: 18463683
- Johanson HC, Chen W, Wicking C, Sturm RA. Inheritance of a novel mutated allele of the OCA2 gene associated with high incidence of oculocutaneous albinism in a Polynesian community. J Hum Genet 2010; 55: 103-11.PMID: 20019752
- Lee ST, Nicholls RD, Bundey S, Laxova R, Musarella M, Spritz RA. Mutations of the P gene in oculocutaneous albinism, ocular albinism, and Prader-Willi syndrome plus albinism. N Engl J Med 1994; 330: 529-34.PMID: 8302318
- Li HY, Wei HY, Zheng H, Qing WR, Duan HL, Meng S, Jiang WY. Prenatal diagnosis of oculocutaneous albinism type II and novel mutations in two Chinese families. Prenat Diagn. 2007; 27: 502-6. PMID: 17385796
- Preising MN, Forster H, Tan H, Lorenz B, de Jong PT, Plomp AS. Mutation analysis in a family with oculocutaneous albinism manifesting in the same generation of three branches. Mol Vis 2007; 13: 1851-5.PMID: 17960121
- Puri N, Gardner JM, Brilliant MH. Aberrant pH of melanosomes in pink-eyed dilution (p) mutant melanocytes. J Invest Dermatol 2000; 115: 607-13. PMID: 10998131
- Rinchik EM, Bultman SJ, Horsthemke B, Lee ST, Strunk KM, Spritz RA, Avidano KM, Jong MT, Nicholls RD. A gene for the mouse pink-eyed dilution locus and for human type II oculocutaneous albinism. Nature 1993; 361: 72-6. PMID: 8421497
- Rooryck C, Morice-Picard F, Elcioglu NH, Lacombe D, Taieb A, Arveiler B. Molecular diagnosis of oculocutaneous albinism: new mutations in the OCA1-4 genes and practical aspects. Pigment Cell Melanoma Res. 2008; 21: 583-7. PMID: 18821858
- Sitaram A, Piccirillo R, Palmisano I, Harper DC, Dell'Angelica EC, Schiaffino MV, Marks MS. Localization to mature melanosomes by virtue of cytoplasmic dileucine motifs is required for human OCA2 function. Mol Biol Cell 2009; 20: 1464-77. PMID: 19116314
- Staleva L, Manga P, Orlow SJ. Pink-eyed dilution protein modulates arsenic sensitivity and intracellular glutathione metabolism. Mol Biol Cell 2002; 13: 4206-20. PMID: 12475946
- Suzuki T, Tomita Y. Recent advances in genetic analyses of oculocutaneous albinism types 2 and 4. J Dermatol Sci. 2008; 51: 1-9. PMID: 18407468
- Toyofuku K, Valencia JC, Kushimoto T, Costin GE, Virador VM, Vieira WD, Ferrans VJ, Hearing VJ. The etiology of oculocutaneous albinism (OCA) type II: the pink protein modulates the processing and transport of tyrosinase. Pigment Cell Res 2002; 15: 217-24. PMID: 12028586
- Wattanasirichaigoon D, Suwannarat P, Thongpradit S. Two novel splice mutations of P gene in a Thai-Chinese patient with oculocutaneous albinism type II (OCA2). J Dermatol Sci 2008; 49: 98-101. PMID: 18036783
- Wei A, Wang Y, Long Y, Wang Y, Guo X, Zhou Z, Zhu W, Liu J, Bian X, Lian S, Li W. A comprehensive analysis reveals mutational spectra and common alleles in chinese patients with oculocutaneous albinism. J Invest Dermatol 2010; 130: 716-24. PMID: 19865097
EDIT HISTORY:
Created by Wei Li & Jonathan Bourne: 07/27/2004
Updated by Wei Li: 04/06/2006
Updated by Wei Li: 08/09/2007
Updated by Wei Li: 02/28/2008
Updated by Wei Li: 08/12/2008
Updated by Wei Li: 02/18/2009
Updated by Wei Li: 03/15/2010